


PRE-IMPLANTATION GENETIC DIAGNOSIS
USING POLAR BODIES
Tomi D1,*, Voigt R1, Eckhold J1, Hinrichs F1, Griesinger G2,
Shulze-Mosgau A2, Schöpper B2, Al-Hasani S2, Diedrich K2, Schwinger E1
*Corresponding Author: Dr. Diana Tomi, Institute of Human Genetics, Medical University of Lübeck, Ratze¬burger Allee 160, 23538 Lübeck, Germany; Tel.: +49(0)451-500-2621; Fax: +49(0)451-500-4187; E-mail: dianatomi@hotmail.com
page: 17
|
|
DISCUSSION
Molecular Genetic Diagnosis for Cystic Fibrosis. Blastomere analysis is the usual and most reliable method of PGD for CF [10]. In Germany, this method is not allowed due to the embryo protection law [2,3]. To establish the possibility of enabling PGD in Germany we developed the analysis on first PBs. To achieve a higher accuracy, the application of sequential analysis of first and second PBs should be done [11,12].
In the first couple (A), three out of five first PBs were heterozygous for the mutation allele (#1, #2 and #4) (Table 1). In PBs #1 and #2, only the wild type allele of the DF508 mutation region was shown. Due to the linked polymorphic markers the detection of heterozygosity of PBs #1 and #2 was possible, even if in the majority of cases only one of the expected alleles was amplified. Polar body #4 showed heterozygosity in four markers. Polar bodies #3 and #5 showed the wild type allele. In PB #5, the result was clearly confirmed by the linked polymorphic markers. In PB #3, two polymorphic markers failed to amplify any allele and two (D7S486 and the T-polythymidine tract) showed alleles linked to the wild type allele. We concluded that the corresponding oocytes carried the mutated allele. The heterozygosity of PB #3 may also be a possible assumption, but this could only be verified by investigation of the second PB. Because three of the five PBs were heterozygous and two carried wild type alleles, no oocyte was deemed appropriate for implantation. Furthermore, none of the oocytes were fertilized.
Allele drop-out (ADO) frequency depends on the cell types analyzed [13], tested genes [14], lysis conditions [15,16] or PCR conditions [17]. In the case of couple A (Table 1), the ADO rate seemed to be high in comparison to earlier published data. After PCR techniques for the second couple were improved and the T-polythymidine tract was replaced by the D7S1842 linked polymorphic marker, it was possible to reduce the ADO rate. In some PBs (for example, #1, #2 and #3 of couple A and #9 of couple B; Table 2), more than two alleles failed to amplify or showed ADO. Since the same technique for PB analysis was done in these cases, ADO may be caused by fragmentation or poor quality of the PBs. In couple B, PB #12 and second PBs #2 and #12 (Table 3) failed to amplify to any of the tested markers. The possible cause for the PCR failure may be due to loss of the PB during transfer or insufficient lysis of the cell coat and nuclear proteins.
Multiplex PCR in single cells showed a similar reliability and ADO rates as singleplex PCR [18]. Amplifying the mutation and linked polymorphic markers in a multiplex PCR assay is a reliable and applicable method in comparison with simple mutation detection in PGD [19,20]. Multiplex PCR, followed by nested PCR, increases accuracy in PBs investigation [7,11]. Moreover, DNA analysis of first and second PBs could increase the accuracy of the diagnosis even more [7,11,21].
In couple B, 12 first PBs were available. Polar body #3 showed the wild type allele and linked polymorphic markers, leading to the conclusion that the oocyte was a carrier of the DF508 mutation. Polar body #11 was heterozygous for the mutation region but all markers showed alleles linked to the mutation. We concluded that the heterozygosity was probably due to a contamination of this PB with the wild type allele. Only oocyte #11 was transferred but no pregnancy was established.
Nine of the first PBs of couple B were heterozygous (Table 2). Strom et al. [21] found approximately 50% of first PBs to be heterozygous for CF. In our case, a higher rate of heterozygosity (9/11; 80%) was found. This finding shows how important the examination of the DNA of the second PB is if no blastomeres are available for PGD.
Some of the second PBs were available in this cycle (Table 3). Two of these were hemizygous (PBs #5 and #10). Polar body #5 showed only the wild type allele and linked polymorphic markers. In PB #10, linked polymorphic markers to the mutation allele were detected but failed to amplify the DF508 mutation itself. In the case of oocytes #5 and #10 a clear diagnosis was possible after examination of the second PB, even in the case of a heterozygous first PB. Second PBs #3 and #4 were heterozygous. Looking at the data of the first PBs (Table 2), the second PB #3 was expected to carry the mutation allele. A possible explanation for its heterozygosity may be the predivision of the chromatids prior to the first meiotic cleavage and a non-disjunction distributing one chromatid carrying the wild type allele in the first PB, while three chromatids remained in the oocyte. After the second meiotic cleavage, two chromatids with the wild type and the mutation allele were distributed in the second PB. A mutation allele probably remained in the oocyte. A nullisomy of chromosome 7 in the oocyte could not be excluded. Angell [22] described a premature division of the centromeres at meiosis I forming single chromatids in oocytes. These findings were confirmed in further investigations and showed the existence of both mechanisms of non-disjunction due to extra chromosomes or extra chromatids [23-25].
Second PB #4 and first PB #4 were heterozygous. The heterozygosity of the first PB is not surprising. A hemizygous second PB with either the mutation allele or the wild type allele was anticipated. Due to the heterozygosity in the second PB we determined a non-disjunction in the second meiotic division of the corresponding oocyte. Aneuploidy as a result of non-disjuction in the second meiotic division is a well known mechanism in some cases of trisomy 18 and trisomy 21 [26,27], but little is known about non-disjunction of chromosome 7 due to rare findings of trisomy 7 in spontaneous abortions. Therefore, interest to investigate this chromosome in PBs or blastomeres for aneuploidy is low. Analyzing PBs and corresponding oocytes by FISH, aneuploidies like disomies and nullisomies of the examined chromosomes 13, 18, 21 and X in first PBs, were found [28].
Several steps were established to avoid contamination: for example, using gowns and gloves, working in different laboratories for pre- and post-PCR, using the equipment for pre-PCR like tubes, racks and pipettes in the laboratory only for single cells and PBs and using blanks for every PCR step. Despite this, contamination occurred in four nested PCR assays (Tables 2 and 3). First PB #11 showed a heterozygosity for the mutation region, while the polymorphic marker showed alleles linked to the mutation region. In this case, contamination of the probe with the wild type allele of the person who handled the PB is feasible. In three other cases (first PBs #3, #4 and second PB #4), the unexpected alleles were not derived from persons who handled the PBs, neither during the ICSI procedure nor in the molecular genetic laboratory.
Molecular Cytogenetic Diagnosis for Aneuploidies. So far, molecular cytogenetic diagnosis for aneuploidies has been performed in only 13 cases due to the recent start of this investigation. Polar body testing for aneuploidies was only offered to couples undergoing ICSI due to male infertility. Aneuploidies of chromosomes 15, 16 and 22 are frequent in abortions [29]. The aim of our study is to increase pregnancy rates by excluding these aneuploidies. Aneuploidy screening in PBs was first reported by Verlinsky et al. [31] using FISH probes for chromosomes X, 18 and/or 13/21. Further investigations with the aim of preventing age related aneuploidies were also performed by Verlinsky et al. [31].
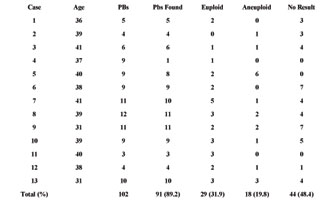
In our study, only the first PBs were tested. In some cases, no diagnosis of the PB was possible due to difficulties in finding PBs on the slide or hybridization failure (Table 4). Ninety-one out of 102 PBs were found on the slides. In one case (#4), a very high rate of loss of PBs was seen due to the initial phase of the investigation. A hybridization failure was detected in 48.4%. These data show that the technique must be improved to achieve a better hybridization rate. Of the tested PBs, 31.9% were euploid for chromosomes 15, 16 and 22. Aneuploidies for further chromosomes cannot be excluded by this method. Due to the short time-span prescribed by the embryo protection law, only a limited amount of chromosomes can be analyzed.
An aneuploidy rate of 19.8% was found by analyzing only first PBs for three chromosomes. Verlinsky et al. [31] found a rate of 43% of aneuploid oocytes analyzing both PBs for three chromosomes. In these cases, the women were older than 35 years. Montag et al. [32] found, in 41.2% of analyzed first PBs, aneuploidies of the chromosomes 13, 16, 18, 21 and 22; the women were between 36 and 40 years old.
Future investigations may increase the hybridization rate. Adding the detection of chromosomes 13 and 18 to the aneuploidy screening, the rate of detected aneuploidies may be increased. In trisomy 16 [33], 15 and 22 [34], non-disjunction of the chromosomes usually occurs in the first meiotic division. So far, the pregnancy rates were not increased when analyzing only the chromosomes 15, 16 and 22, so that it will be important to add analysis of chromosomes 13, 18 and 21. For this analysis, a kit by Vysis can be used for chromosomes 13, 16, 18, 21 and 22. Approximately 50% of trisomy 18 and 30% of trisomy 13 are developed in the second meiotic division. For more accurate results, examination of the second PB will be necessary.
Table 1. Polar bodies of couple A. The alleles of the mother are shown in the last row. Red alleles are linked to the mutation, green alleles to the wild type allele. (T-Poly = T-polythymidine tract.)

Table 2. First polar bodies of couple B. The alleles of the mother are shown in the last row. Red alleles are linked to the mutation, green alleles to the wild type allele. Contamination is indicated by an exclamation mark.
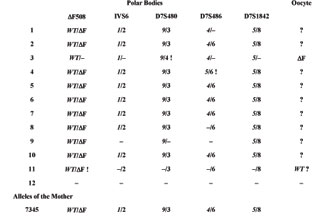
Table 3. Second polar bodies of couple B. The alleles of the mother are showed in the lowest line. Red alleles are linked to the mutation, green alleles to the wild type allele. Contamination is indicated by an exclamation mark.

Table 4. Thirteen cases of polar body diagnosis for aneuploidies. Euploid, aneuploid and polar bodies with no results are shown in relation to the number of polar bodies found on the slides.
|
|
|
|



 |
Number 27
VOL. 27 (2), 2024 |
Number 27
VOL. 27 (1), 2024 |
Number 26
Number 26 VOL. 26(2), 2023 All in one |
Number 26
VOL. 26(2), 2023 |
Number 26
VOL. 26, 2023 Supplement |
Number 26
VOL. 26(1), 2023 |
Number 25
VOL. 25(2), 2022 |
Number 25
VOL. 25 (1), 2022 |
Number 24
VOL. 24(2), 2021 |
Number 24
VOL. 24(1), 2021 |
Number 23
VOL. 23(2), 2020 |
Number 22
VOL. 22(2), 2019 |
Number 22
VOL. 22(1), 2019 |
Number 22
VOL. 22, 2019 Supplement |
Number 21
VOL. 21(2), 2018 |
Number 21
VOL. 21 (1), 2018 |
Number 21
VOL. 21, 2018 Supplement |
Number 20
VOL. 20 (2), 2017 |
Number 20
VOL. 20 (1), 2017 |
Number 19
VOL. 19 (2), 2016 |
Number 19
VOL. 19 (1), 2016 |
Number 18
VOL. 18 (2), 2015 |
Number 18
VOL. 18 (1), 2015 |
Number 17
VOL. 17 (2), 2014 |
Number 17
VOL. 17 (1), 2014 |
Number 16
VOL. 16 (2), 2013 |
Number 16
VOL. 16 (1), 2013 |
Number 15
VOL. 15 (2), 2012 |
Number 15
VOL. 15, 2012 Supplement |
Number 15
Vol. 15 (1), 2012 |
Number 14
14 - Vol. 14 (2), 2011 |
Number 14
The 9th Balkan Congress of Medical Genetics |
Number 14
14 - Vol. 14 (1), 2011 |
Number 13
Vol. 13 (2), 2010 |
Number 13
Vol.13 (1), 2010 |
Number 12
Vol.12 (2), 2009 |
Number 12
Vol.12 (1), 2009 |
Number 11
Vol.11 (2),2008 |
Number 11
Vol.11 (1),2008 |
Number 10
Vol.10 (2), 2007 |
Number 10
10 (1),2007 |
Number 9
1&2, 2006 |
Number 9
3&4, 2006 |
Number 8
1&2, 2005 |
Number 8
3&4, 2004 |
Number 7
1&2, 2004 |
Number 6
3&4, 2003 |
Number 6
1&2, 2003 |
Number 5
3&4, 2002 |
Number 5
1&2, 2002 |
Number 4
Vol.3 (4), 2000 |
Number 4
Vol.2 (4), 1999 |
Number 4
Vol.1 (4), 1998 |
Number 4
3&4, 2001 |
Number 4
1&2, 2001 |
Number 3
Vol.3 (3), 2000 |
Number 3
Vol.2 (3), 1999 |
Number 3
Vol.1 (3), 1998 |
Number 2
Vol.3(2), 2000 |
Number 2
Vol.1 (2), 1998 |
Number 2
Vol.2 (2), 1999 |
Number 1
Vol.3 (1), 2000 |
Number 1
Vol.2 (1), 1999 |
Number 1
Vol.1 (1), 1998 |
|
|

